Last Monday, I offered a dark scenario to start off your workweek: the prospect of a highly pathogenic zoonotic virus spreading feverishly around the globe—or, as I should have titled the column: “Hi, everyone. How was your weekend?”
Today, I am determined to keep things upbeat—so again, let’s discuss the ongoing threat of pandemic. But this time, here’s a far more hopeful spin: It may well be that the oldest threat to humankind—the spread of infection—is the one where the newest digital health technology can have the greatest impact.
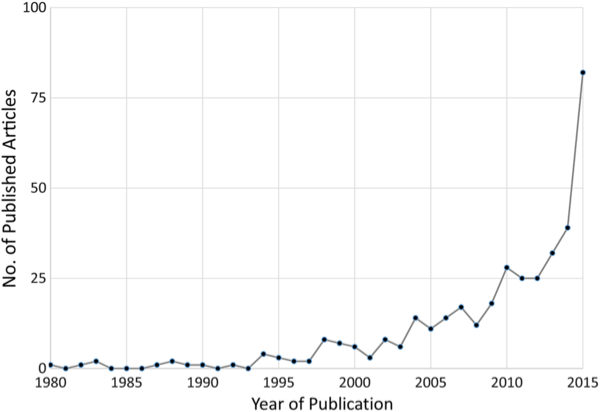
The Journal of Infectious Diseases, the flagship publication of the Infectious Diseases Society of America, devoted its December issue to ways in which big data—and a host of new electronic tracking platforms for both medical and non-medical events—can be used to vastly improve our traditional disease surveillance systems (which, frankly, are terribly flawed now). And with the number of scholarly articles on the subject having risen exponentially since 2001 (see the chart below), there’s a tremendous amount of information out there about what works and what doesn’t.
From: Big Data for Infectious Disease Surveillance and Modeling
As Jihye Choi and colleagues at Seoul National University’s School of Public Health reviewed in another journal this past December (really worth reading), there are three fundamental limitations of many current disease-surveillance systems. The first is language interoperability. Most tend to be in English—even when the centers of outbreaks are frequently in non-English-speaking regions. Second, the networks are often “excessively complex” requiring costly upkeep. And finally, these largely passive systems, run by national public health authorities, often lag in reporting local events—a big problem when speed is of the essence. (That’s what we saw, for example, in the latest Ebola outbreak in West Africa.)
Two new approaches worth checking out are ResistanceOpen, developed by a team of epidemiologists, clinicians, and data experts at Boston Children’s Hospital, and the Global Virome Project.
The first is a very cool interactive global map tracking reports of antimicrobial resistance, based on both public records and data submitted by independent users of this “open-collaboration database.” (Want a cheap thrill? Type in your zip code here and discover what superbugs are lurking in your backyard.)
The second is a more comprehensive ten-year project—proposed by a world-spanning alliance of experts—to preempt emerging infectious disease threats by characterizing virtually all zoonotic viruses that have epidemic potential—and creating a real-time, open-access database to monitor them. It’s an ambitious—and worthwhile—endeavor. (You can read more about it here.)
This essay appears in today’s edition of the Fortune Brainstorm Health Daily. Get it delivered straight to your inbox.










